This article is co-authored with generative AI. While I have cross-checked facts against official documentation where possible, errors may remain. Please verify primary sources before making important decisions.
The Kouigenjimonogatari Text DB is a database that publishes the text of Kouigenjimonogatari — a critical edition of The Tale of Genji compiled by Kikan Ikeda — as TEI/XML. The base scans are taken from the National Diet Library Digital Collections, where the work is available as out-of-copyright. The site includes a statistics page that visualizes per-chapter pages, lines, characters, and waka (Japanese poems) counts.
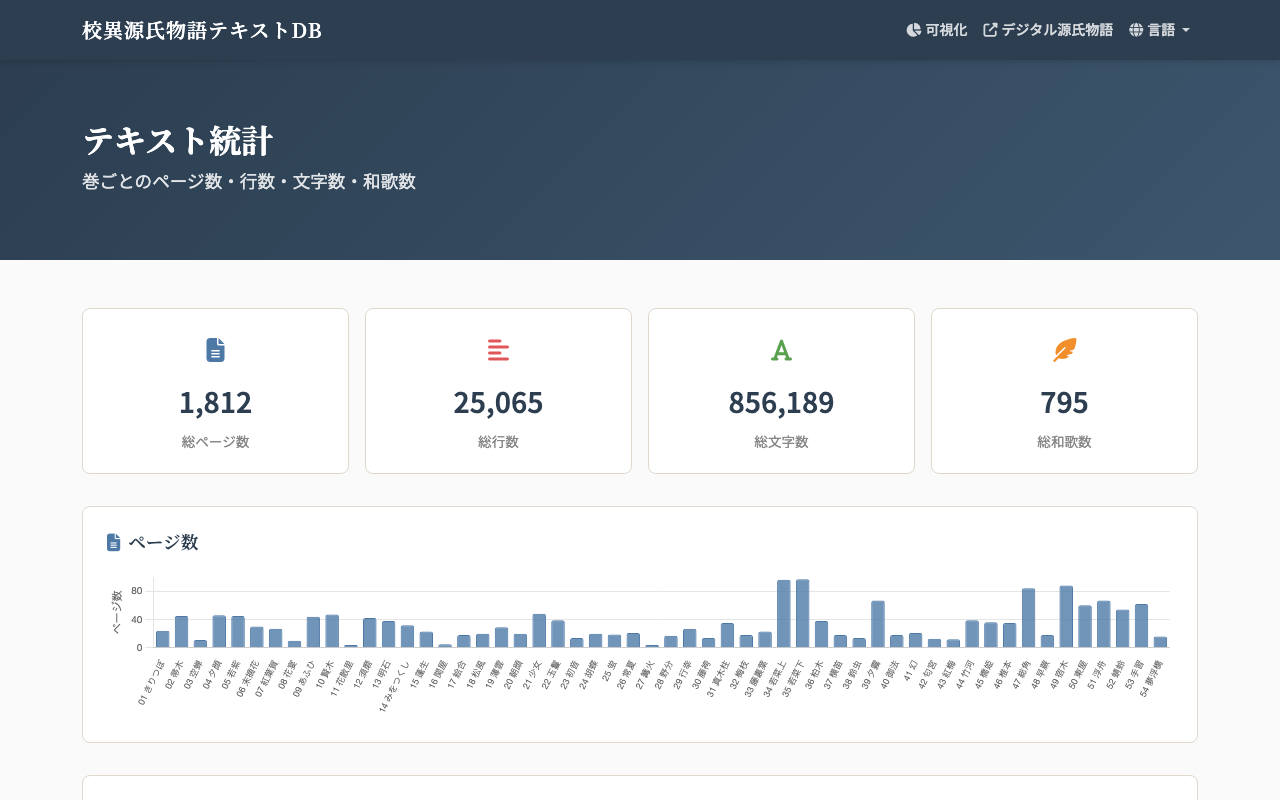
This post records how such a statistics page is generated from TEI/XML-structured transcription data, and how its rebuild and redeployment are automated via GitHub Actions.
What the statistics page shows
https://kouigenjimonogatari.github.io/stats.html shows:
- Four summary cards: total pages, total lines, total characters, and total waka.
- Per-chapter bar charts (Chart.js) for pages, lines, characters, and waka.
At the time of writing, all 54 chapters have been transcribed. The total character count is 856,189 across 1,812 pages. Distribution skews are also visible — for example, Wakana-jō and Wakana-ge are markedly longer than the other chapters, with Wakana-ge being the single longest chapter.
Structuring transcription text in TEI/XML
TEI (Text Encoding Initiative) is an international guideline set for encoding humanities texts. Plain-text files make it hard to determine programmatically where one page ends and the next begins, or which lines form a waka. Once the same content is marked up with TEI-conformant XML tags, downstream aggregation and conversion become much easier.
In the Kouigenjimonogatari Text DB, each chapter is encoded roughly like this:
<TEI xmlns="http://www.tei-c.org/ns/1.0">
<teiHeader>
<!-- Metadata: bibliography, license, contributors -->
</teiHeader>
<facsimile>
<!-- Bindings to IIIF images at the National Diet Library Digital Collections -->
</facsimile>
<text>
<body>
<p>
<pb n="5" facs="..."/> <!-- Page break: page 5 -->
<lb/>
<seg corresp="..."> <!-- Line / clause unit -->
いつれの御時にか...
</seg>
<lb/>
<seg corresp="..."> <!-- waka is nested inside seg as lg -->
<lg xml:id="waka-001" type="waka" rhyme="tanka">
<l n="1">...</l>
<l n="2">...</l>
<l n="3">...</l>
<l n="4">...</l>
<l n="5">...</l>
</lg>
</seg>
</p>
</body>
</text>
</TEI>
Because elements like <pb> (page break), <seg> (line / clause), and <lg type="waka"> (waka group) make their meaning explicit, it is straightforward to count “how many pages?”, “how many lines?”, and “how many waka?” programmatically.
The aggregation script
The repository’s scripts/prebuild.py includes a function that aggregates per-chapter statistics into a JSON file:
def build_chapters_json():
"""Aggregate pages, lines, characters, and waka per chapter, then write chapters.json."""
chapters = []
for ch in CHAPTERS: # ['01', '02', ..., '54']
src_path = os.path.join(XML_LW_DIR, f'{ch}.xml')
if not os.path.exists(src_path):
continue
with open(src_path, 'r', encoding='utf-8') as f:
content = f.read()
# Number of <pb> elements = page count
pages = len(re.findall(r'<pb\s', content))
# Number of <seg> elements = line count; sum of inner text = character count
seg_texts = re.findall(r'<seg[^>]*>(.*?)</seg>', content)
lines = len(seg_texts)
chars = sum(
len(re.sub(r'<[^>]+>', '', seg).strip())
for seg in seg_texts
)
# Number of <lg type="waka"> elements = waka count
waka = len(re.findall(r'<lg\s[^>]*type="waka"', content))
chapters.append({
'chapter': ch,
'chapter_name': CHAPTER_NAMES.get(ch, ch), # e.g. '01' -> 'Kiritsubo'
'pages': pages,
'lines': lines,
'chars': chars,
'waka': waka,
})
with open('docs/data/chapters.json', 'w', encoding='utf-8') as f:
json.dump(chapters, f, ensure_ascii=False, indent=2)
Regular expressions are sufficient here precisely because TEI makes structural meaning explicit through tags. Doing the same aggregation against plain text would force more brittle, heuristic-based judgments — for example, deciding what counts as a chapter boundary or how to identify a waka.
Rendering on the static page
docs/stats.html is a plain static page that fetches the generated docs/data/chapters.json and uses Chart.js to draw bar charts:
fetch('data/chapters.json')
.then(r => r.json())
.then(chapters => {
const totalPages = chapters.reduce((s, c) => s + c.pages, 0);
const totalChars = chapters.reduce((s, c) => s + c.chars, 0);
// Update the four summary cards
document.getElementById('totalPages').textContent = totalPages.toLocaleString();
document.getElementById('totalChars').textContent = totalChars.toLocaleString();
// Draw the bar chart
new Chart(document.getElementById('pagesChart'), {
type: 'bar',
data: {
labels: chapters.map(c => c.chapter_name),
datasets: [{ data: chapters.map(c => c.pages), label: 'Pages' }],
},
});
// Lines, characters, and waka follow the same pattern
});
The page is plain HTML + JavaScript, so it needs no server-side runtime or database. Static hosting such as GitHub Pages is enough.
CI/CD: rebuild and redeploy on push
The site is published on GitHub Pages, and updates to the TEI/XML propagate to the live page within a few minutes. The pipeline is configured in .github/workflows/deploy.yml. The relevant excerpt is:
on:
push:
branches: [master]
paths:
- 'xml/master/**'
- 'scripts/**'
- 'docs/**'
- 'data/**'
- 'requirements.txt'
- 'package.json'
jobs:
build:
runs-on: ubuntu-latest
steps:
- uses: actions/checkout@v4
- uses: actions/setup-python@v5
with: { python-version: '3.12' }
- run: pip install -r requirements.txt
- uses: actions/setup-java@v4
with: { distribution: 'temurin', java-version: '21' }
# Download Saxon-HE and xmlresolver (used by XSLT 3.0)
- run: |
curl -sL -o /tmp/saxon-he.jar \
https://repo1.maven.org/maven2/net/sf/saxon/Saxon-HE/12.5/Saxon-HE-12.5.jar
curl -sL -o /tmp/xmlresolver.jar \
https://repo1.maven.org/maven2/org/xmlresolver/xmlresolver/6.0.4/xmlresolver-6.0.4.jar
- name: Run prebuild
env:
SAXON_JAR: /tmp/saxon-he.jar:/tmp/xmlresolver.jar
run: python3 scripts/prebuild.py tei xsl waka stats epub
- uses: actions/upload-pages-artifact@v3
with: { path: docs }
deploy:
needs: build
runs-on: ubuntu-latest
environment:
name: github-pages
steps:
- uses: actions/deploy-pages@v4
The main steps are:
- The workflow runs whenever
xml/master/**,scripts/**,docs/**, etc. are updated. - It sets up Python and Java, then places Saxon-HE (an XSLT 3.0 processor).
prebuild.pygenerates TEI-derived files (display TEI, HTML, waka list, statistics, EPUB, etc.).- The
docs/directory is uploaded as a Pages artifact and deployed viaactions/deploy-pages.
Because docs/data/chapters.json is regenerated in the stats step of prebuild.py, any update to the TEI/XML transcription is reflected automatically on the statistics page. There is no manual number-rewriting step in the workflow.
Why structuring text as TEI/XML pays off
This is a more general observation than the statistics page, but encoding meaning in TEI/XML lets you generate multiple downstream artifacts from the same source. The current site ships the following derived outputs:
- The statistics page (the topic of this post)
- Vertically-typeset HTML for reading (via XSLT)
- A waka list (extracted from
<lg type="waka">and serialized as JSON) - EPUB (per chapter)
- PDF (XSLT to TeX, then typeset with lualatex)
- Linked Data (RDF) and a SPARQL endpoint (browsed via the SNORQL UI bundled with the repository)
- A DTS (Distributed Text Services) API-compatible viewer (the external service
dts-typescript.vercel.appreads this site’s TEI/XML)
The more derived outputs there are, the more the upfront cost of TEI encoding is amortized. With plain text, similar pre-processing tends to be reinvented for each downstream target.
A statistics page like https://kouigenjimonogatari.github.io/stats.html emerges from a simple flow: text is structured as TEI/XML, the build step aggregates and writes JSON, the static HTML and JavaScript fetch and render the JSON, and CI/CD rebuilds and redeploys on push. Reducing the implementation cost of these derived pages is one of the practical reasons to adopt TEI/XML for transcription data.